Neutrophils (or polymorphonuclear leukocytes, PMNs) are the second most abundant cell type after red blood cells in the blood and make up to 40-70% of the leukocytes in adults.1 Their main immune functions include phagocytosis, degranulation, and formation of neutrophil extracellular traps (NETs). They also secrete several cytokines and chemokines that regulate inflammation.2 Neutrophils pose several challenges in vitro due their short half-life of 6-8 hours and extreme sensitivity to apoptosis and activation. In addition, they do not proliferate well and are hard to expand or cryopreserve. Moreover, isolating specific neutrophil populations is challenging with classical methods, such as density gradients since the method can increase polarization responses that results in the induction of artificial activation of neutrophils.3,4
Due to these challenges in studying neutrophils, a microfluidic cell sorter, e.g. the WOLF G2, is well suited by allowing the user to select specific neutrophil populations and perform gentle, low-activation sorts to ensure better viability and purity.
Neutrophil Isolation
Sample preparation was performed at room temperature. Whole blood was processed within two hours of collection and neutrophils were isolated using a double gradient method. Blood was diluted 1:1 with a buffer composed of HBSS (Invitrogen, #14025092), 1% BSA (Invitrogen, #37525), and 2mM EDTA (Invitrogen, #AM9260G) that was layered over a double gradient composed of Leuko Spin Medium (pluriSelect, #60-0091-10) and PBMC Spin Medium (pluriSelect, #60- 00092-10) into a 15mL conical tube ensuring that the gradients and blood do not fully mix. The sample tube was centrifuged at 1000 x g for 30 minutes with no brake. This resulted in a separated granulocyte layer that was transferred into a new tube. The granulocytes were washed with HBSS/1% BSA/2mM EDTA and centrifuged at 300 x g for 10 minutes. The supernatant was removed and the cells were then resuspended with 500μL of sorting buffer that was composed of RPMI (ATCC, #30-2001), 0.5% BSA, and 5mM EDTA to then lyse the red blood cells using 10mL of 1X RBC Lysis Buffer (BioLegend, #420301). The cells were then washed with HBSS, 1% BSA, and 2mM EDTA. Finally, the cells were resuspended with sorting buffer and diluted to a concentration of 1.0 x 107 cells/ mL for staining and control preparations.
Neutrophil Staining for Sorting with the WOLF
The isolated granulocytes were blocked for non-specific binding with Human TruStain FcX (Fc Receptor Blocking Solution) (BioLegend, #422302) and True-Stain Monocyte Blocker (BioLegend, #426102) and incubated at room temperature for 15 minutes.
After blocking, the sample was stained with CD45 APC-Cy7 (BioLegend, #304014), CD16 AF488 (BioLegend, #302022), CD11b PE (BioLegend, #301406), and SYTOX AADvanced (Invitrogen, #R37168). The sample was diluted to have a final concentration of 3 x 105 cells/mL and filtered with a 37 μm filter cap prior to analysis and sorting with the WOLF. In addition, a sample was prepared and stimulated with a cell activation cocktail (BioLegend, #423301) to function as a control for neutrophil activation. Moreover, a compensation matrix was created using single-stained controls, SpectraComp Compensation Beads (Slingshot Bio, #SSB-05-A), and ViaComp Beads (Slingshot Bio, #SSBS-003).
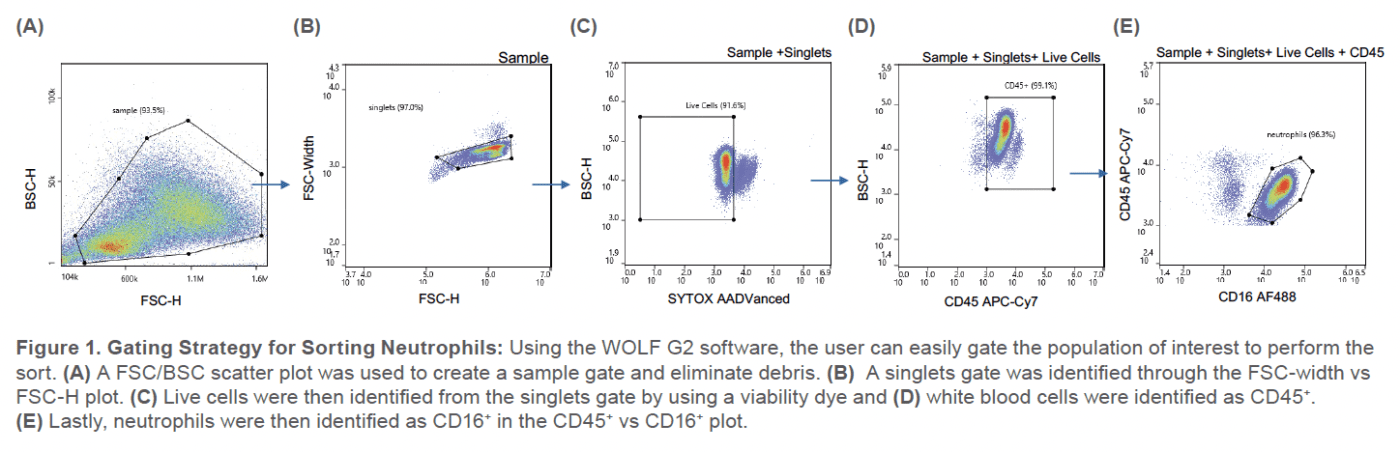
The population of interest consisted of live neutrophils that were CD16-AF488 positive and SYTOX AADvanced negative (Figure 1). 1 x 105 cells were sorted into a collection tube containing sorting buffer.
Post-Sort Analysis
The sorted sample was incubated at 37°C, 5% CO2 for two hours and then reanalyzed with the WOLF. Approximately 5 x 103 of the previously sorted and incubated cells were analyzed for viability (SYTOX AADvanced negative), purity (CD16-AF488 positive), and neutrophil activation based on the MFI of CD11b+ cells.
This work was repeated with two technical replicates for each of the two donors (biological replicates).
Purity and Viability
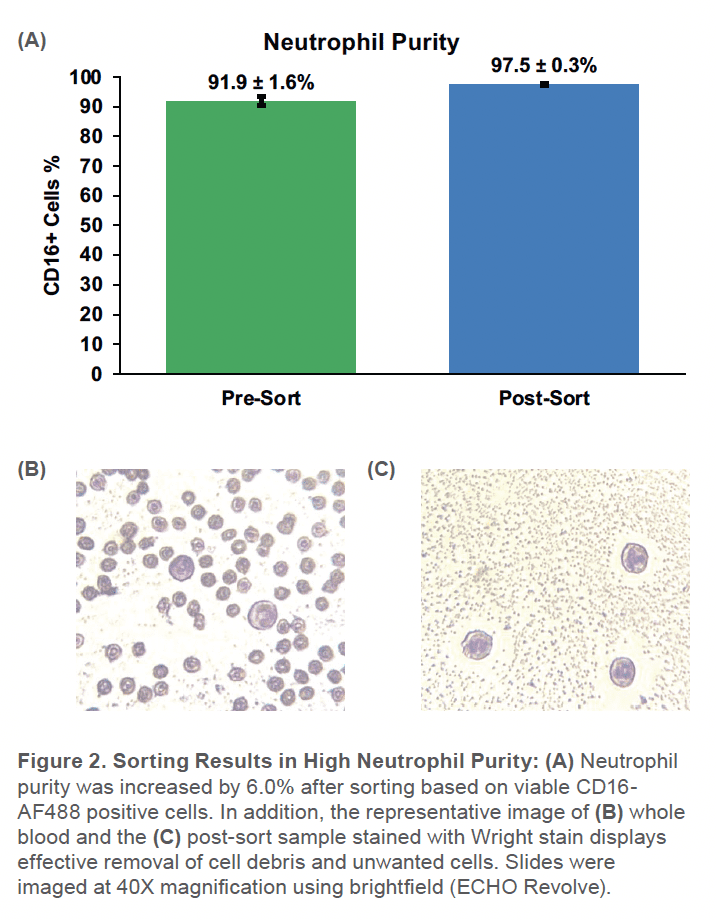
When sorting with the WOLF, the purity increased from 92 ± 1.9% SD to 98 ± 0.3% SD (Figure 2). Additionally, the viability went from 88 ± 1.6%SD to 97 ± 0.1%SD (Figure 3). Although starting with a gradient-enriched sample, the WOLF further removed dead neutrophils that can continue releasing cytokines and increase the number of activated neutrophils in the sample.5
Neutrophil Activation
No change in the number of activated neutrophils was seen in the sorted neutrophils, relative to unsorted controls (Figure 4). In contrast, the activated control showed a substantial twofold increase in the number of CD11b+ activated neutrophils compared to the post-sort sample (Figure 4).
By using the WOLF G2 for neutrophil isolation, researchers can perform gentle sorts with minimal neutrophil activation, higher viability, and better purity to support downstream applications or to discover new emerging therapeutics
For more information, visit nanocellect.com or email [email protected]
APN-034





